Non-Coding RNA May Serve as Blood-based Biomarker for ALS
Written by |
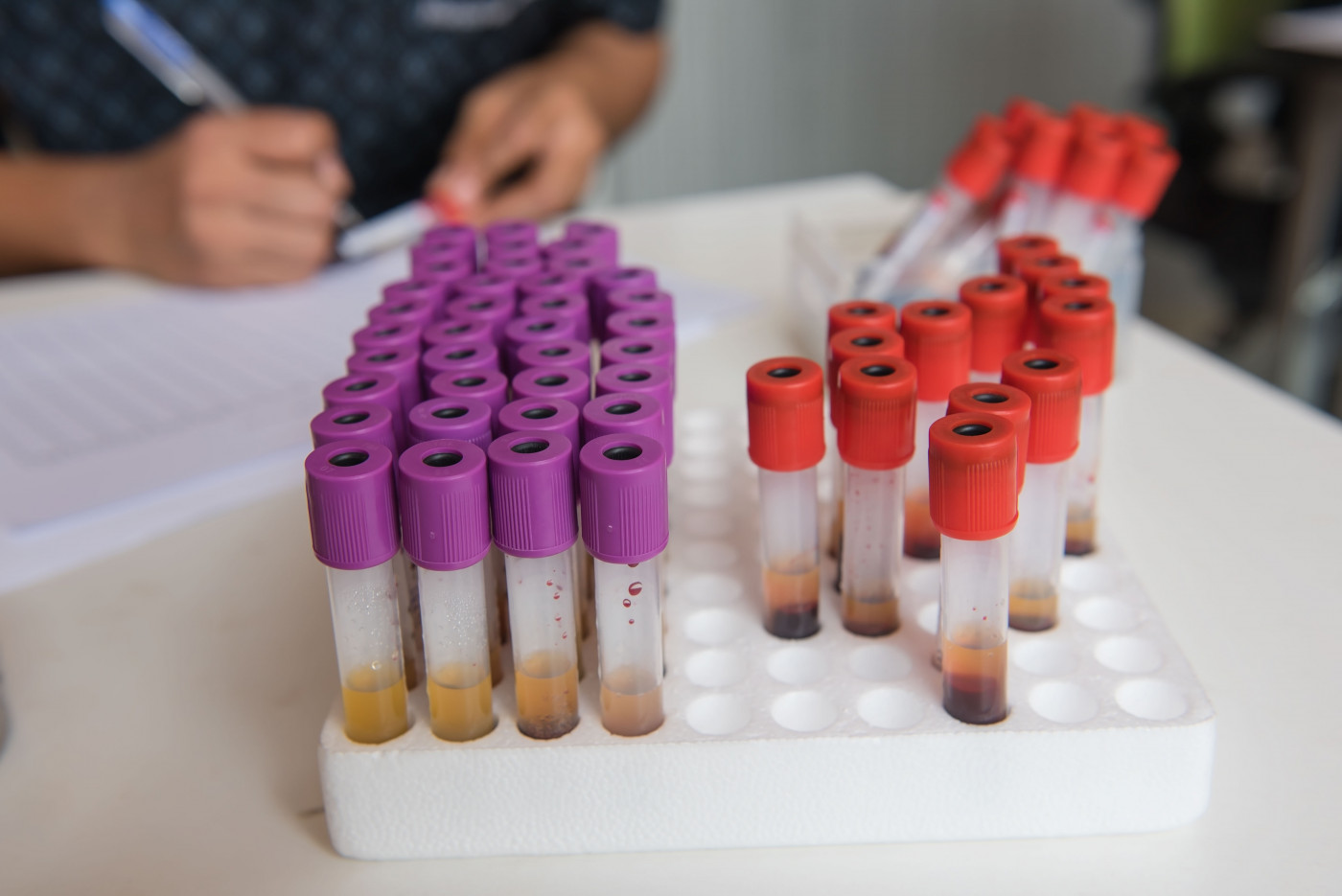
Distinct RNA patterns in a person’s blood may signal the presence of amyotrophic lateral sclerosis (ALS), a study reports.
Researchers, led by Majid Hafezparast, PhD, a professor of molecular neuroscience at the University of Sussex in the U.K., discovered that certain types of non-coding RNA (ncRNA), a kind of nucleic acid, differed among people with ALS, those with similar disorders, and healthy individuals.
These results could improve the process of diagnosing ALS and help detect it earlier in patients. Today, an ALS diagnosis can take up to a year.
“In order to effectively diagnose and treat ALS, we are in urgent need of biomarkers as a tool for early diagnosis and for monitoring the efficacy of therapeutic interventions in clinical trials,” Hafezparast said in a press release.
The study, “Identification of a potential non-coding RNA biomarker signature for amyotrophic lateral sclerosis,” was published in the journal Brain Communications.
Non-coding RNA have become attractive targets in the search for ALS biomarkers. RNA is often known as an intermediary between DNA and proteins. The DNA that codes for a gene is first transcribed into a strand of RNA, which is then used as an instruction manual of sorts for building a protein.
Some DNA, however, code for RNA that never goes on to become a protein. These ncRNA species carry out a variety of regulatory roles throughout the body, often serving to control the expression of other genes — the process by which information in a gene is synthesized to create a protein.
Hafezparast and his colleagues hypothesized that ncRNA may be dysregulated in ALS, which could potentially be behind much of the variation in clinical ALS signs.
The group analyzed blood samples from 245 patients with ALS and similar disorders and healthy people used as controls, using RNA-seq to look at each one’s ncRNA composition. They found seven ncRNA, whose expression, or the amount found in a sample, appeared markedly different between affected individuals and healthy controls.
The team used the relative amounts of these seven ncRNA to build a computational model that could be used to categorize patients based on their blood sample. This model accurately categorized more than 90% of their subjects as either ALS patients (included both slow- and fast-progressing patients) or non-ALS controls (including healthy controls, neurological controls, and disease mimics).
Because models tend to perform best when tested on the samples that were used to train them, the group confirmed their results in a separate group of patients. The algorithm was able to correctly identify these ALS and non-ALS control subjects nearly 74% of the time.
Lending credence to the team’s results, one ncRNA found in the ALS signature, called hsa-miR-206, has been previously linked to ALS, both biologically and as a biomarker. This RNA, known as a microRNA, or miRNA, occurs most frequently in muscles and may be released into the bloodstream as a result of muscle death.
While these results demonstrate the strength of using RNA-seq to identify and measure small ncRNA, the researchers noted that low amounts of intact small RNA and high amounts of degraded longer RNA in the blood continue to pose a challenge to the development of accurate blood-based biomarker tests.
Further studies will need to test whether the biomarkers seen here are present in individuals before symptom onset, which would be vital to early diagnosis and to any therapies aimed at preventing the onset of symptoms to begin with.
Future studies will also need to evaluate the usefulness of these biomarkers in prognosis, or disease progression.
“This is an important initial step in the analysis of all species of ncRNA in the serum for establishing a biomarker signature for ALS and highlights the potential of this approach for other neurodegenerative diseases,” the researchers concluded.



Leave a comment
Fill in the required fields to post. Your email address will not be published.